import numpy as np
np.random.seed(42)
import matplotlib.pyplot as plt
from scipy.special import gamma as gamma_fn
from pyapprox.util.backends.numpy import NumpyBkd
from pyapprox.surrogates.kernels import Matern32Kernel, ExponentialKernel
from pyapprox.surrogates.kle import MeshKLE
from pyapprox.pde.galerkin.mesh.structured import StructuredMesh1D
from pyapprox.pde.galerkin.basis.lagrange import LagrangeBasis
from pyapprox.pde.field_maps.kle_factory import (
create_spde_matern_kle,
create_fem_nystrom_nodes_kle,
)
bkd = NumpyBkd()
rng = np.random.default_rng(7)KLE III: SPDE-Based KLE for Matern Fields
PyApprox Tutorial Library
Learning Objectives
After completing this tutorial, you will be able to:
- Explain the SPDE connection between Matern covariance kernels and differential operators
- Use
create_spde_matern_kleto build a sparse SPDE-based KLE - Compare SPDE eigenvalues against the dense kernel-based
MeshKLE - Understand boundary artefacts and how they diminish as the domain grows
- Choose between
MeshKLE,DataDrivenKLE, andSPDEMaternKLEfor a given problem
In KLE I we introduced the KLE and in KLE II we compared kernel-based and data-driven approaches. Both rely on dense \(O(N^2)\) matrices. The SPDE approach replaces the dense covariance with a sparse PDE operator whose Green’s function is the Matern covariance, giving \(O(N)\) memory.
The Matern-SPDE connection
A Matern random field with smoothness \(\nu\) and correlation length \(\ell_c\) is the solution to the SPDE:
\[(\kappa^2 - \Delta)^{\alpha/2} u = \mathcal{W}\]
with \(\alpha = \nu + d/2\) (integer \(\alpha\) avoids fractional operators). For the bilaplacian prior \(\alpha = 2\):
| Dimension \(d\) | Smoothness \(\nu = 2 - d/2\) | Kernel |
|---|---|---|
| 1D | \(\nu = 3/2\) | Matern-3/2 |
| 2D | \(\nu = 1\) | Between Matern-1/2 and 3/2 |
| 3D | \(\nu = 1/2\) | Exponential (Matern-1/2) |
The SPDE parameters map to physical parameters as: \[\gamma \leftrightarrow \text{diffusion}, \quad \delta = \kappa^2 \gamma, \quad \ell_c = \sqrt{\gamma/\delta} = 1/\kappa\]
from scipy.special import kv
from pyapprox.tutorial_figures._kle import plot_matern_kernels_paths
x_plot = np.linspace(0, 2, 200)
mesh_plot = bkd.array(x_plot[None, :])
x0 = 1.0
ell = 0.4
# Compute Matern kernel curves
r = np.abs(x_plot - x0)
kernel_curves = []
for nu, name, color in [(0.5, "Matern-1/2\n(exponential)", "steelblue"),
(1.0, "Matern-1 \n(SPDE 2D)", "darkorange"),
(1.5, "Matern-3/2\n(SPDE 1D)", "green"),
(2.5, "Matern-5/2", "red")]:
sqrt2nu = np.sqrt(2*nu)
z = sqrt2nu * np.maximum(r, 1e-14) / ell
k = (2**(1-nu) / gamma_fn(nu)) * z**nu * kv(nu, z)
k[r < 1e-12] = 1.0
kernel_curves.append((r, k, name, color))
# Compute sample paths for two smoothness values
dx_plot = x_plot[1] - x_plot[0]
w_plot = np.ones(200) * dx_plot; w_plot[0] /= 2; w_plot[-1] /= 2
quad_plot = bkd.array(w_plot)
sample_paths_list = []
path_labels = []
for nu, name, kernel_class in [
(0.5, "Matern-1/2", ExponentialKernel),
(1.5, "Matern-3/2", Matern32Kernel),
]:
kern = kernel_class(bkd.array([ell]), (0.01, 10.0), 1, bkd)
kle_plot = MeshKLE(
mesh_plot, kern, sigma=1.0, nterms=50,
quad_weights=quad_plot, bkd=bkd,
)
coef = bkd.array(rng.standard_normal((50, 5)))
sample_paths_list.append(bkd.to_numpy(kle_plot(coef)))
path_labels.append(name)
fig, axes = plt.subplots(1, 3, figsize=(14, 4))
plot_matern_kernels_paths(x_plot, kernel_curves, sample_paths_list, path_labels, axes)
plt.tight_layout()
plt.show()
Smaller \(\nu\) produces rougher (less differentiable) paths. The SPDE approach lets us work with any \(\nu\) through the integer \(\alpha\) parametrization.
Building the SPDE KLE with create_spde_matern_kle
The create_spde_matern_kle factory assembles the sparse FEM precision operator \(A = \gamma K_{\text{stiff}} + \delta M + \xi M_{\partial}\) and solves the generalized eigenvalue problem \(A\phi_k = \mu_k M \phi_k\). The KLE eigenvalues are \(\lambda_k = \gamma^2/(\tau^2 \mu_k^2)\).
This uses only sparse matrices, giving \(O(N)\) memory instead of \(O(N^2)\).
from pyapprox.tutorial_figures._kle import plot_spde_overview
# Parameters for ell_c = 0.5 in 1D
ell_c = 0.5
gamma = 1.0
delta = gamma / ell_c**2
nterms = 6
nx = 300
# Build FEM mesh and basis
mesh = StructuredMesh1D(nx=nx, bounds=(0.0, 2.0), bkd=bkd)
basis = LagrangeBasis(mesh, degree=1)
# Create SPDE KLE
kle_spde = create_spde_matern_kle(
basis, n_modes=nterms, gamma=gamma, delta=delta,
sigma=1.0, bkd=bkd,
)
lam_spde = bkd.to_numpy(kle_spde.eigenvalues())
phi_spde = bkd.to_numpy(kle_spde.eigenvectors())
x_nodes = bkd.to_numpy(basis.dof_coordinates())[0]
# Compare with dense Matern-3/2 MeshKLE
nu = 1.5
kappa = np.sqrt(delta / gamma)
rho = np.sqrt(2 * nu) / kappa
lenscale = bkd.array([rho])
matern_kernel = Matern32Kernel(lenscale, (0.01, 100.0), 1, bkd)
skfem_basis = basis.skfem_basis()
kle_nystrom = create_fem_nystrom_nodes_kle(
skfem_basis, matern_kernel, nterms=nterms, sigma=1.0, bkd=bkd,
)
lam_dense = bkd.to_numpy(kle_nystrom.eigenvalues())
nsamples_show = 5
Z_s = bkd.array(rng.standard_normal((nterms, nsamples_show)))
fields_spde = bkd.to_numpy(kle_spde(Z_s))
var_spde = bkd.to_numpy(kle_spde.pointwise_variance())
k_idx = np.arange(1, nterms + 1)
fig, axes = plt.subplots(1, 3, figsize=(14, 4))
plot_spde_overview(
x_nodes, phi_spde, nterms, fields_spde, var_spde,
k_idx, lam_dense, lam_spde, ell_c,
axes[0], axes[1], axes[2],
)
plt.tight_layout()
plt.show()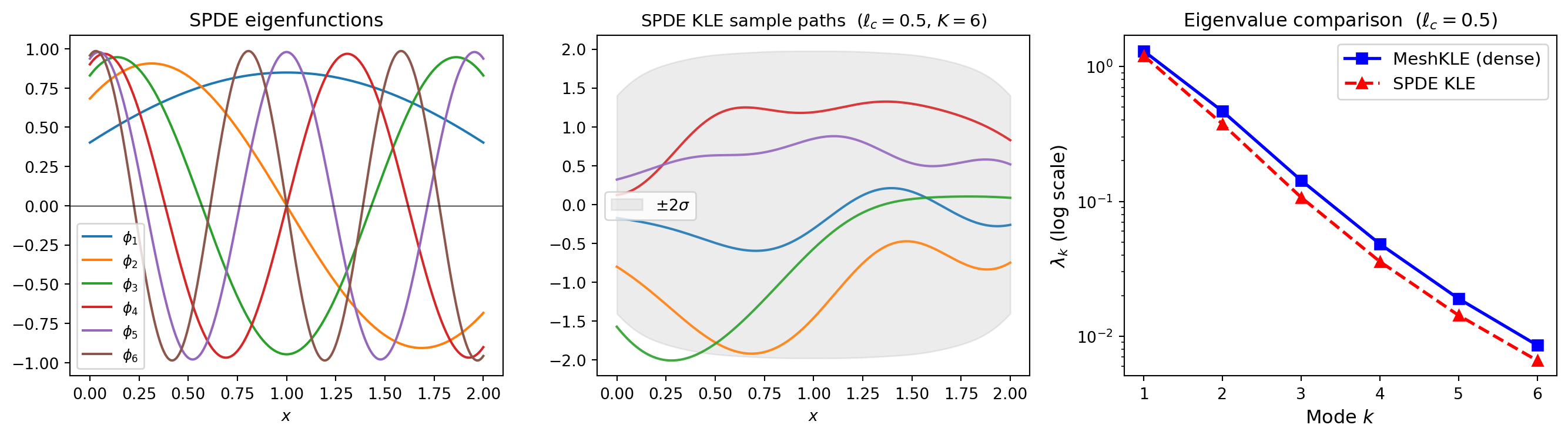
pointwise_variance(). Right: eigenvalue comparison between the sparse SPDE KLE and the dense MeshKLE with Matern-3/2 kernel. They agree at low modes but differ near the boundary.
The SPDE and dense eigenvalues agree at low modes. They differ near the boundary – an intrinsic feature of the SPDE approach.
Boundary effects: the key SPDE limitation
The Matern covariance on \(\mathbb{R}^d\) is stationary (translation invariant). On a bounded domain the SPDE uses Robin boundary conditions that modify the covariance near the edges. This boundary artefact decreases as the domain grows relative to \(\ell_c\).
from pyapprox.tutorial_figures._kle import plot_boundary_artefacts
ell_c_fixed = 0.5
domain_data = []
for domain_len in [1.0, 2.0, 5.0]:
nx_d = max(100, int(domain_len / 0.01))
mesh_d = StructuredMesh1D(nx=nx_d, bounds=(0.0, domain_len), bkd=bkd)
basis_d = LagrangeBasis(mesh_d, degree=1)
gamma_d = 1.0
delta_d = gamma_d / ell_c_fixed**2
nterms_d = min(40, nx_d - 2)
kle_d = create_spde_matern_kle(
basis_d, n_modes=nterms_d, gamma=gamma_d, delta=delta_d,
sigma=1.0, bkd=bkd,
)
x_d = bkd.to_numpy(basis_d.dof_coordinates())[0]
var_d = bkd.to_numpy(kle_d.pointwise_variance())
domain_data.append((x_d, var_d, domain_len))
fig, axes = plt.subplots(1, 3, figsize=(14, 4))
plot_boundary_artefacts(domain_data, ell_c_fixed, axes)
fig.suptitle("Boundary artefact decreases as domain grows relative to $\\ell_c$",
fontsize=12)
plt.tight_layout()
plt.show()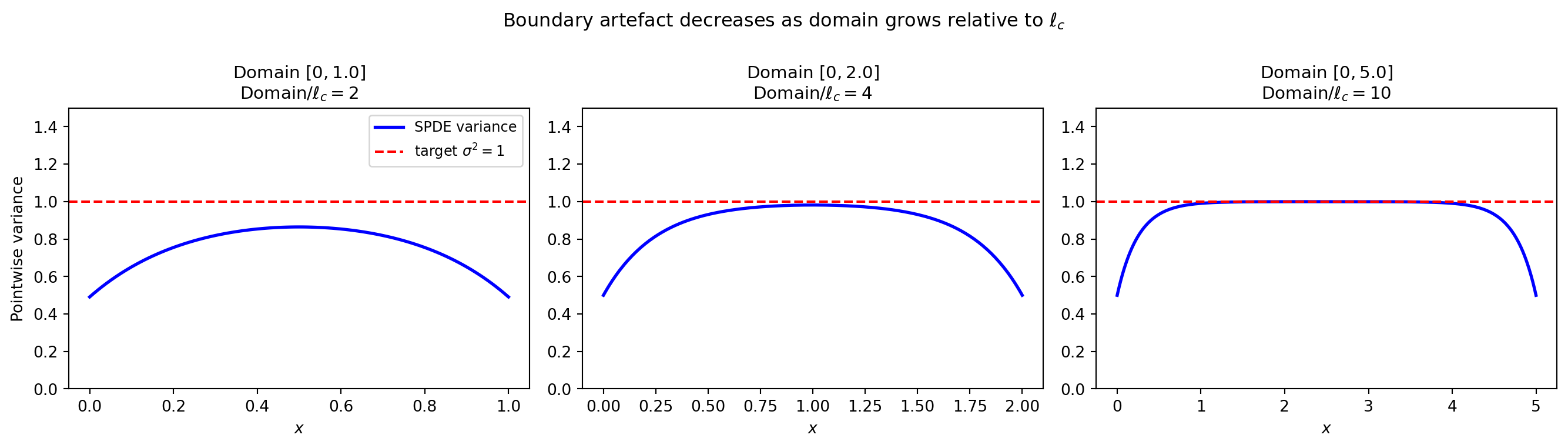
Rule of thumb: for reliable interior statistics extend the domain by \(\gtrsim 3\ell_c\) on each side, then discard the boundary buffer.
Memory and computational scaling
The eigenvalue solve cost also differs significantly:
MeshKLE: Dense eigendecomposition costs \(O(N^2 K)\) (or \(O(N^3)\) for all modes)DataDrivenKLE: SVD of the \(N \times N_s\) sample matrix costs \(O(N \cdot N_s \cdot K)\)SPDEMaternKLE: Sparse generalized eigenvalue solve costs \(O(N \cdot K)\) with iterative methods (e.g. ARPACK)
from pyapprox.tutorial_figures._kle import plot_memory_scaling
# Scaling data
N_vals = np.array([100, 300, 500, 1000, 2000, 5000])
K_modes = 20
mem_dense = N_vals**2 * 8 / 1e6
mem_sparse = N_vals * 3 * 8 / 1e6
cost_dense = N_vals**2 * K_modes
cost_sparse = N_vals * K_modes
# Sample paths with different correlation lengths
ells = [0.3, 0.5, 0.8]
nx_s = 200
mesh_s = StructuredMesh1D(nx=nx_s, bounds=(0.0, 2.0), bkd=bkd)
basis_s = LagrangeBasis(mesh_s, degree=1)
x_s = bkd.to_numpy(basis_s.dof_coordinates())[0]
fields_by_ell = []
for ell_target in ells:
gamma_t = 1.0
delta_t = gamma_t / ell_target**2
kle_t = create_spde_matern_kle(
basis_s, n_modes=20, gamma=gamma_t, delta=delta_t,
sigma=1.0, bkd=bkd,
)
Z_t = bkd.array(rng.standard_normal((20, 3)))
fields_by_ell.append(bkd.to_numpy(kle_t(Z_t)))
fig, axes = plt.subplots(1, 2, figsize=(12, 4))
plot_memory_scaling(
N_vals, mem_dense, mem_sparse, cost_dense, cost_sparse,
x_s, fields_by_ell, ells, axes[0], axes[1],
)
plt.tight_layout()
plt.show()
For \(N = 5000\) nodes the dense matrix requires 200 MB vs 0.12 MB for the sparse SPDE operator. In 2D with \(N = 100 \times 100 = 10^4\) nodes the difference is 800 MB vs 0.24 MB – the SPDE approach becomes the only feasible option.
Choosing \(\gamma\), \(\delta\), \(\xi\) from physical parameters
Given target correlation length \(\ell_c\) and marginal std \(\sigma\):
from pyapprox.tutorial_figures._kle import plot_spde_parameters
# Empirical covariance at midpoint for each correlation length
mid = nx_s // 2
cov_by_ell = []
for ell_target in ells:
gamma_t = 1.0
delta_t = gamma_t / ell_target**2
kle_t = create_spde_matern_kle(
basis_s, n_modes=20, gamma=gamma_t, delta=delta_t,
sigma=1.0, bkd=bkd,
)
nmc = 2000
Z_mc = bkd.array(rng.standard_normal((20, nmc)))
F = bkd.to_numpy(kle_t(Z_mc))
cov_by_ell.append(F[mid, :] @ F.T / nmc)
# Convergence of SPDE toward dense KLE as domain grows
domain_sizes = [5, 10, 20, 40]
n_modes_conv = 5
max_errors = []
gamma_conv, delta_conv, sigma_conv = 1.0, 1.0, 1.0
kappa_conv = np.sqrt(delta_conv / gamma_conv)
rho_conv = np.sqrt(2 * 1.5) / kappa_conv
lenscale_conv = bkd.array([rho_conv])
kernel_conv = Matern32Kernel(lenscale_conv, (0.01, 500.0), 1, bkd)
for L in domain_sizes:
nx_conv = 10 * L
mesh_conv = StructuredMesh1D(nx=nx_conv, bounds=(0.0, float(L)), bkd=bkd)
basis_conv = LagrangeBasis(mesh_conv, degree=1)
kle_spde_conv = create_spde_matern_kle(
basis_conv, n_modes=n_modes_conv, gamma=gamma_conv, delta=delta_conv,
sigma=sigma_conv, bkd=bkd,
)
kle_nystrom_conv = create_fem_nystrom_nodes_kle(
basis_conv.skfem_basis(), kernel_conv, nterms=n_modes_conv,
sigma=sigma_conv, bkd=bkd,
)
eig_spde = bkd.to_numpy(kle_spde_conv.eigenvalues())
eig_nystrom = bkd.to_numpy(kle_nystrom_conv.eigenvalues())
rel_err = np.abs(eig_spde - eig_nystrom) / eig_nystrom
max_errors.append(np.max(rel_err))
fig, axes = plt.subplots(1, 2, figsize=(12, 4))
plot_spde_parameters(
domain_sizes, max_errors, x_s, cov_by_ell, ells, mid,
axes[0], axes[1],
)
plt.tight_layout()
plt.show()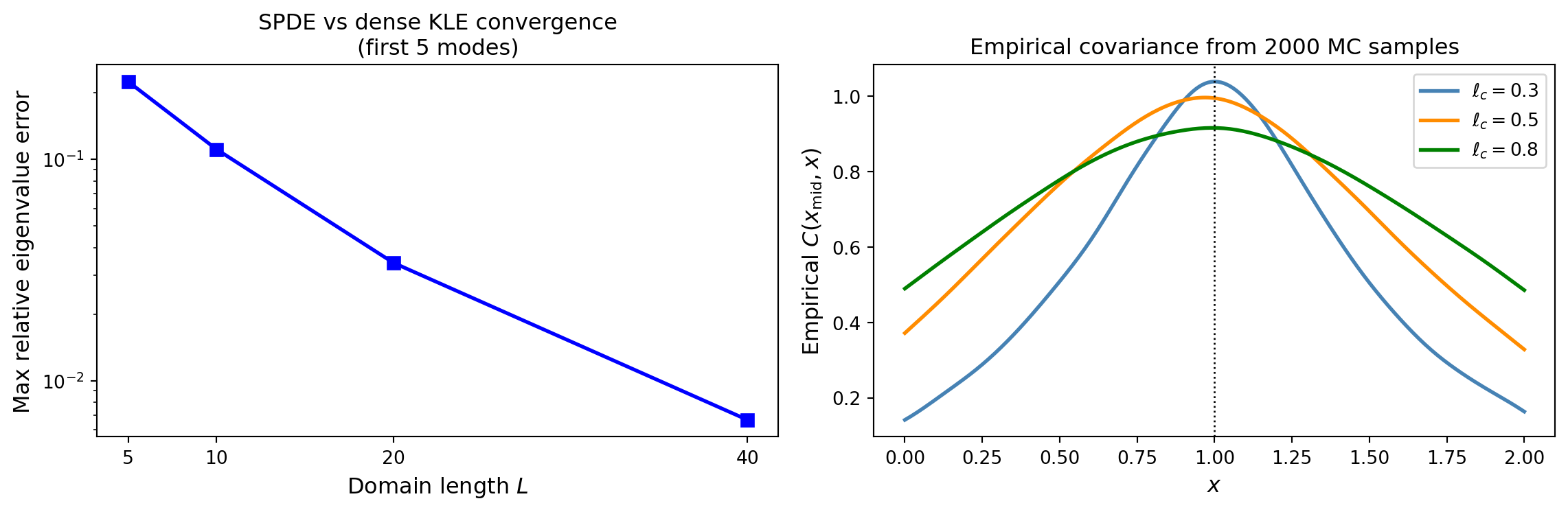
Summary: which KLE method to use?
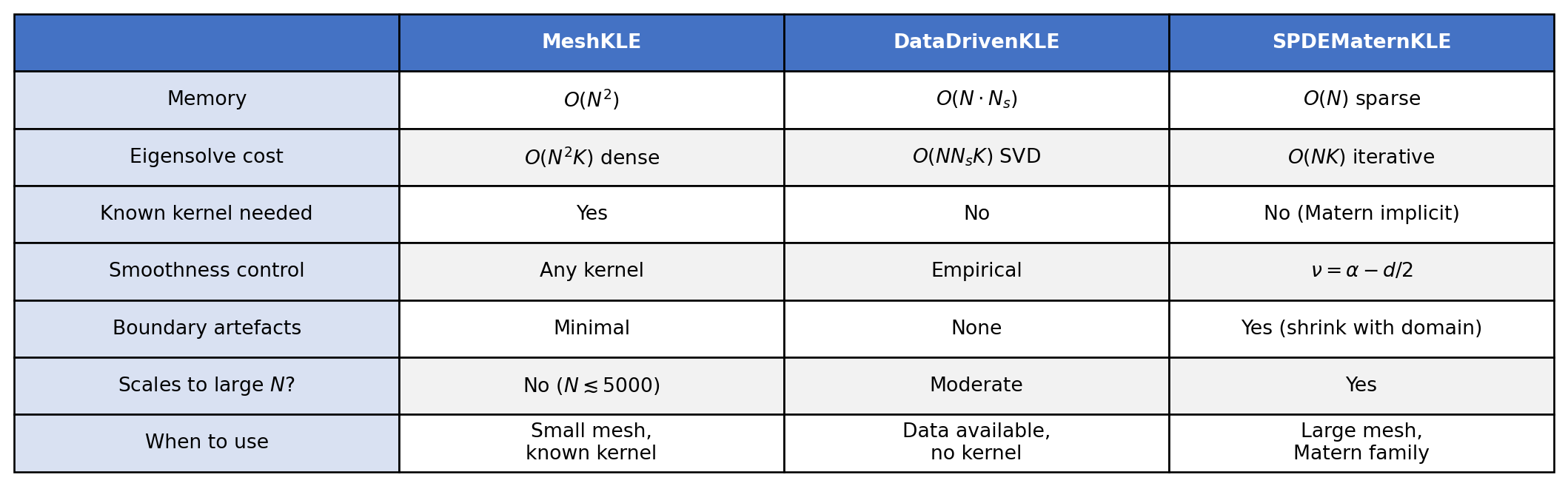
MeshKLE (dense kernel), DataDrivenKLE (SVD of samples), and SPDEMaternKLE (sparse PDE operator).