import numpy as np
import matplotlib.pyplot as plt
from pyapprox.util.backends.numpy import NumpyBkd
from pyapprox.expdesign.benchmarks import LinearGaussianOEDBenchmark
from pyapprox.expdesign.diagnostics import KLOEDDiagnostics
np.random.seed(1)
bkd = NumpyBkd()Bayesian OED: Verifying Convergence with KLOEDDiagnostics
PyApprox Tutorial Library
Learning Objectives
After completing this tutorial, you will be able to:
- Construct a
LinearGaussianOEDBenchmarkand query its exact EIG - Use
KLOEDDiagnosticsto decompose total MSE into squared bias and variance - Confirm empirically that outer sample count \(M\) drives variance while inner count \(N_\text{in}\) drives bias
- Read convergence rates from
compute_convergence_rateand match them to theory
Prerequisites
Complete The Double-Loop Estimator before this tutorial. The estimator has two sample budgets: outer (\(M\)) controls variance; inner (\(N_\text{in}\)) controls the bias from the evidence approximation. We verify both empirically here.
Setup
# Problem parameters — match test_kl_oed_convergence.py for reproducibility
nobs = 5
degree = 2
noise_std = 0.5
prior_std = 0.5
benchmark = LinearGaussianOEDBenchmark(nobs, degree, noise_std, prior_std, bkd)
diagnostics = KLOEDDiagnostics(benchmark)
# Uniform design: equal weight on every candidate observation
weights = bkd.ones((nobs, 1)) / nobsExact EIG as Ground Truth
LinearGaussianOEDBenchmark provides a closed-form EIG for any design weights, giving us an exact reference against which to measure numerical error.
exact = diagnostics.exact_eig(weights)A single numerical estimate with moderate sample counts:
approx = diagnostics.compute_numerical_eig(
nouter=200,
ninner=100,
design_weights=weights,
seed=42,
)compute_numerical_eig is fully reproducible when called with the same seed. Always record your seed alongside results so experiments can be re-run exactly.
MSE Decomposition: Bias and Variance
compute_mse runs nrealizations independent estimates and decomposes total MSE into squared bias and variance: \(\text{MSE} = \text{bias}^2 + \text{variance}\).
bias, variance, mse = diagnostics.compute_mse(
nouter=100,
ninner=50,
nrealizations=20,
design_weights=weights,
base_seed=0,
)Outer vs. Inner: Which Budget Matters More?
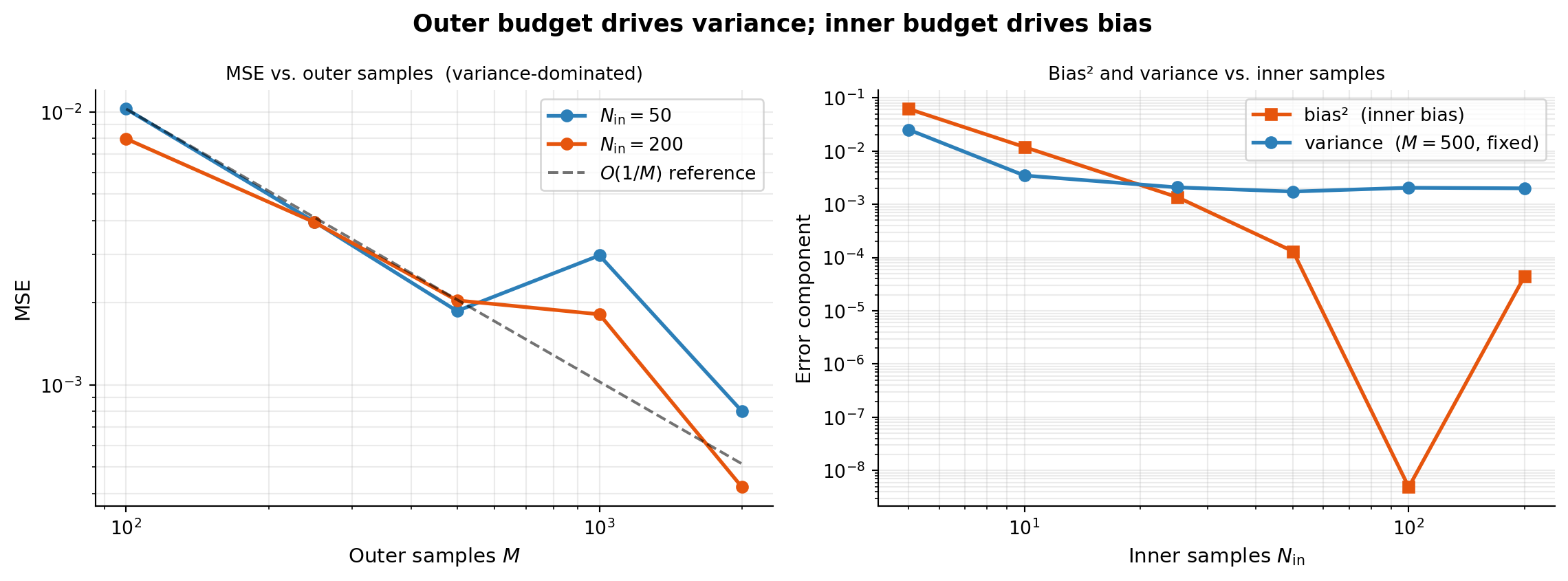
The two figures below answer this directly. The left panel sweeps \(M\) (outer samples) with fixed \(N_\text{in}\), showing how MSE decays with the outer budget. The right panel sweeps \(N_\text{in}\) (inner samples) with fixed \(M\), showing how bias² and variance each respond to the inner budget.
outer_counts = [100, 250, 500, 1000, 2000]
inner_counts = [50, 200]
M_fixed = 500
inner_sweep = [5, 10, 25, 50, 100, 200]
# Outer sweep
values_mc = diagnostics.compute_mse_for_sample_combinations(
outer_sample_counts=outer_counts,
inner_sample_counts=inner_counts,
nrealizations=30,
design_weights=weights,
base_seed=0,
)
# Inner sweep: fix M, vary N_in
values_inner = diagnostics.compute_mse_for_sample_combinations(
outer_sample_counts=[M_fixed],
inner_sample_counts=inner_sweep,
nrealizations=30,
design_weights=weights,
base_seed=0,
)
# Convergence rates
for ii, ninner in enumerate(inner_counts):
rate = KLOEDDiagnostics.compute_convergence_rate(
outer_counts,
bkd.to_numpy(values_mc["mse"][ii]).tolist(),
)A rate near 1.0 confirms the \(O(1/M)\) variance decay. The right panel shows bias² falling steeply once \(N_\text{in} \gtrsim 50\), after which MSE is dominated by variance from the outer loop. Increasing \(N_\text{in}\) beyond that point yields diminishing returns — invest budget in \(M\) instead.
For faster convergence with the same \(M\), see MC vs. Halton for EIG Estimation.
Key Takeaways
LinearGaussianOEDBenchmarkprovides an exact EIG reference for any weights.KLOEDDiagnostics.compute_msedecomposes total MSE into squared bias and variance across independent realizations.- The outer sample count \(M\) controls variance and follows \(O(1/M)\); the inner count \(N_\text{in}\) controls bias, which becomes negligible once \(N_\text{in} \gtrsim 50\) for this problem.
- Once bias² is small relative to variance, further increasing \(N_\text{in}\) wastes compute — increase \(M\) instead.
Exercises
Run
compute_msewithnouter=50, ninner=5and compare bias² to variance. At this extreme, which term dominates?From the right panel, estimate the convergence rate of bias² with respect to \(N_\text{in}\). Is it faster or slower than the \(O(1/M)\) outer rate? Why?
Try
degree=4(more parameters). Does the exact EIG increase or decrease relative todegree=2? Does the \(N_\text{in}\) needed to suppress bias change?
Next Steps
- MC vs. Halton for EIG Estimation — achieving faster convergence by replacing Monte Carlo with quasi-random sequences
- Goal-Oriented Bayesian OED — when prediction accuracy matters more than parameter estimation