import numpy as np
import matplotlib.pyplot as plt
from pyapprox.util.backends.numpy import NumpyBkd
from pyapprox.expdesign.benchmarks import LinearGaussianPredOEDBenchmark
from pyapprox.expdesign.diagnostics import (
PredictionOEDDiagnostics,
create_prediction_oed_diagnostics,
)
np.random.seed(1)
bkd = NumpyBkd()Goal-Oriented OED: Convergence Verification
PyApprox Tutorial Library
Learning Objectives
After completing this tutorial, you will be able to:
- Construct a
LinearGaussianPredOEDBenchmarkand query its analytical utility - Use
PredictionOEDDiagnosticsto compute MSE, bias, and variance of numerical estimates - Explain why the inner loop (not outer) drives convergence for goal-oriented OED
- Compare MC convergence rates for the standard deviation utility
Prerequisites
Read Goal-Oriented Bayesian OED and Gaussian Posterior Expressions.
Quick Recap
From the analysis tutorial, the expected standard deviation of the posterior push-forward in a linear Gaussian model is:
\[ U_\mathrm{std}(\mathbf{w}) = \sqrt{\boldsymbol{\psi}\boldsymbol{\Sigma}_\star(\mathbf{w})\boldsymbol{\psi}^\top}. \]
This is deterministic — no averaging over observations is needed. The numerical estimator must reproduce this from Monte Carlo samples alone. Because the estimator uses ratios of QoI-weighted likelihoods, the bias from the inner loop dominates, unlike the EIG estimator.
Setup
nobs = 2
degree = 3
noise_std = 0.5
prior_std = 0.5
npred = 1
benchmark = LinearGaussianPredOEDBenchmark(
nobs, degree, noise_std, prior_std, npred, bkd,
)
diagnostics = create_prediction_oed_diagnostics(benchmark, "linear_stdev")
# Uniform design weights
weights = bkd.ones((nobs, 1)) / nobsAnalytical Reference Value
exact = diagnostics.exact_utility(weights)A single numerical estimate:
approx = diagnostics.compute_numerical_utility(
nouter=500,
ninner=1000,
design_weights=weights,
seed=42,
)MSE Decomposition
bias, variance, mse = diagnostics.compute_mse(
nouter=100,
ninner=500,
nrealizations=50,
design_weights=weights,
base_seed=0,
)Compare to the EIG case in boed_kl_usage.qmd: there, outer samples drive the MSE. Here, the inner samples drive it. You can verify this by holding ninner fixed while increasing nouter — the MSE barely changes.
MSE vs. Inner Sample Count (MC)
outer_counts = [100, 500] # fixed at modest values
inner_counts = [100, 500, 1000, 5000]
values_mc = diagnostics.compute_mse_for_sample_combinations(
outer_sample_counts=outer_counts,
inner_sample_counts=inner_counts,
nrealizations=50,
design_weights=weights,
base_seed=0,
)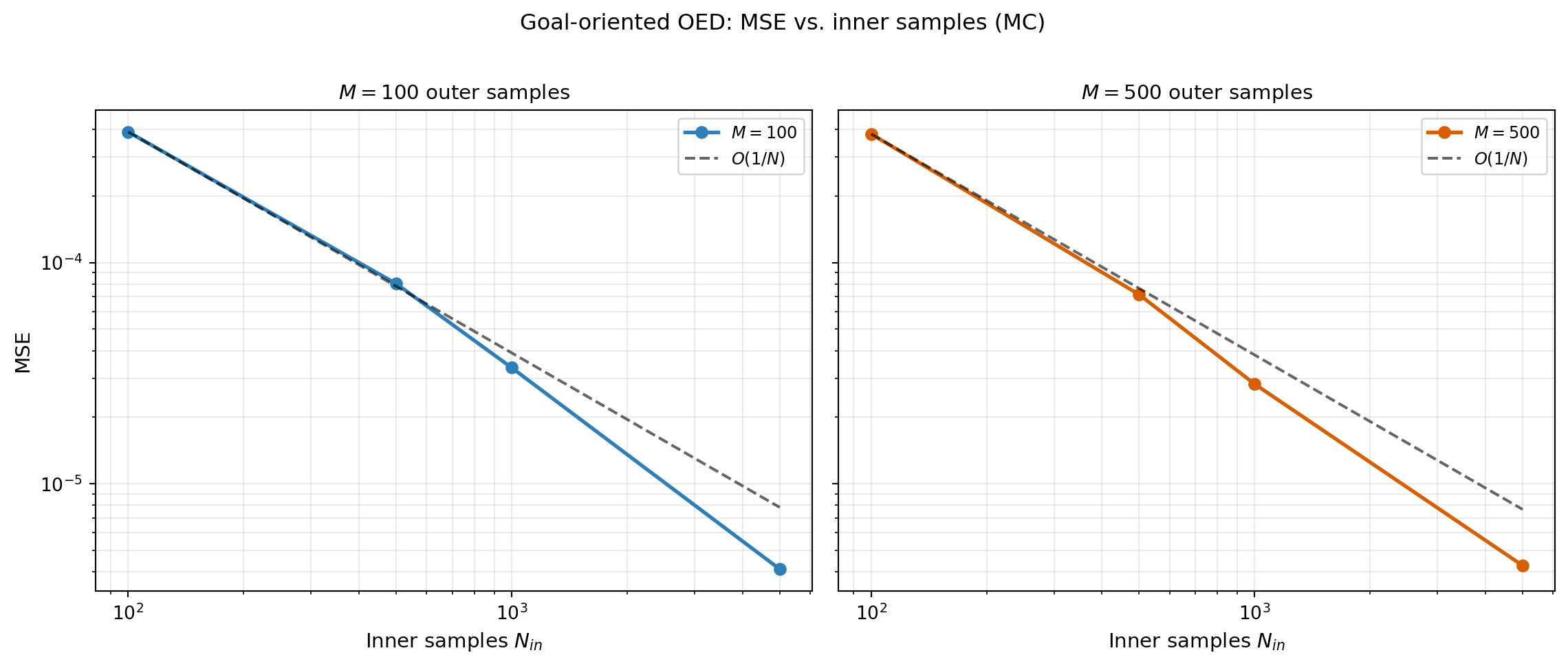
# MC convergence rates (w.r.t. inner samples)
mse_array = bkd.to_numpy(bkd.vstack(values_mc["mse"])) # (n_inner, n_outer)
for jj, nouter in enumerate(outer_counts):
mse_values = mse_array[:, jj].tolist()
rate = PredictionOEDDiagnostics.compute_convergence_rate(inner_counts, mse_values)AVaR Deviation Variant
Switching to AVaR deviation requires only changing the utility type in the create_prediction_oed_diagnostics call.
diagnostics_avar = create_prediction_oed_diagnostics(
benchmark, "linear_avar", beta=0.9,
)
exact_avar = diagnostics_avar.exact_utility(weights)Key Takeaways
- For goal-oriented OED, inner samples dominate MSE convergence — increasing outer samples alone barely reduces error.
PredictionOEDDiagnosticsmirrorsKLOEDDiagnostics: samecompute_mse,compute_mse_for_sample_combinations, andcompute_convergence_rateinterface.- Switching between standard deviation and AVaR utility requires only changing
utility_typein thecreate_prediction_oed_diagnosticscall.
Exercises
Run
compute_mse_for_sample_combinationsvaryingouter_countswithninnerfixed at 2000. Confirm that the MSE does not decrease.Try
npred=3(three QoI components). Does the analytical utility change? Does the convergence rate change?Compare
exact_utilityfor designs that concentrate all weight on one observation vs. uniform weights. Which is better for this polynomial model?
Next Steps
- Optimize the design weights using the gradient expressions from Bayesian OED: Gradients and the Reparameterization Trick adapted for the goal-oriented utility
- Extend to nonlinear models where no analytical reference is available