Parameter Inference with MCMC
This example (ex_minf_sketch.py) walks through a complete Bayesian
parameter inference workflow for a 1-D model using adaptive MCMC. It
covers data generation, inference configuration, MCMC sampling, and
rich post-processing visualisation.
Source: ex_minf_sketch.py
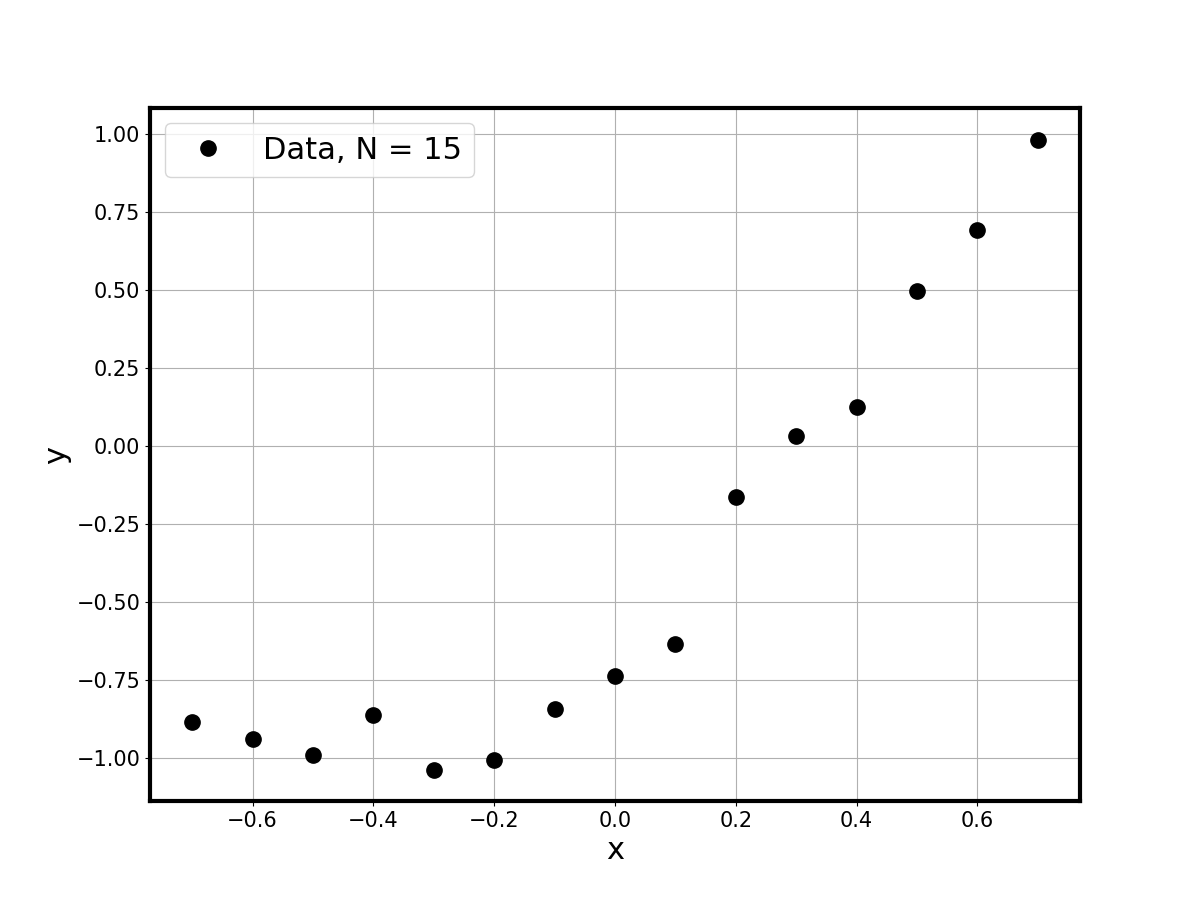
Synthetic Data
A tanh truth model is evaluated on a grid and noisy observations are
generated at 15 equally-spaced points:
true_model = tanh_model # tanh(3*(x - 0.3))
npt = 15
xdata = 0.7 * np.linspace(-1., 1., npt).reshape(-1, 1)
ydata = true_model(xdata) + 0.1 * np.random.randn(npt, 1)
Calibration Model
An exponential model with two parameters is used as the calibration surrogate:
def exp_model_single(par, xx):
return par[1] * np.exp(par[0] * xx) - 2
The model is deliberately simpler than the truth, so the inference must compensate through its posterior uncertainty.
Inference Configuration
The inference is configured through a single dictionary that controls every aspect of the calibration:
model_infer_config = {
"inpdf_type" : 'pci', # PC-based internal PDF
"pc_type" : 'LU', # Legendre-Uniform basis
"outord" : 1, # first-order output PC
"rndind" : [0, 1], # both params are random
"lik_type" : 'gausmarg', # Gaussian marginal likelihood
"pr_type" : 'uniform', # uniform prior (unbounded)
"dv_type" : 'var_fixed', # known data variance
"dv_params" : [0.01], # sigma^2 = 0.1^2
"calib_type" : 'amcmc', # adaptive MCMC
"calib_params": {
'nmcmc': 10000, 't0': 100,
'tadapt': 100, 'gamma': 0.1
},
}
Running MCMC
A single call launches the full inference pipeline — BFGS initialisation followed by adaptive MCMC:
calib_results = minf.model_infer(
ydata, calib_model, xdata, calib_model_pdim,
**model_infer_config)
The chain, log-posterior, and acceptance rates are returned in
calib_results.
Post-Processing
Posterior model samples are generated by evaluating the calibration model at the MCMC chain points and collecting predictive statistics:
checkmode = [calib_results['post'].model, xgrid,
range(len(xgrid)), md_transform]
ycheck_pred, samples = minf.model_infer_postp(
calib_results, checkmode=checkmode,
nburn=len(calib_results['chain']) // 10,
nevery=10, nxi=1)
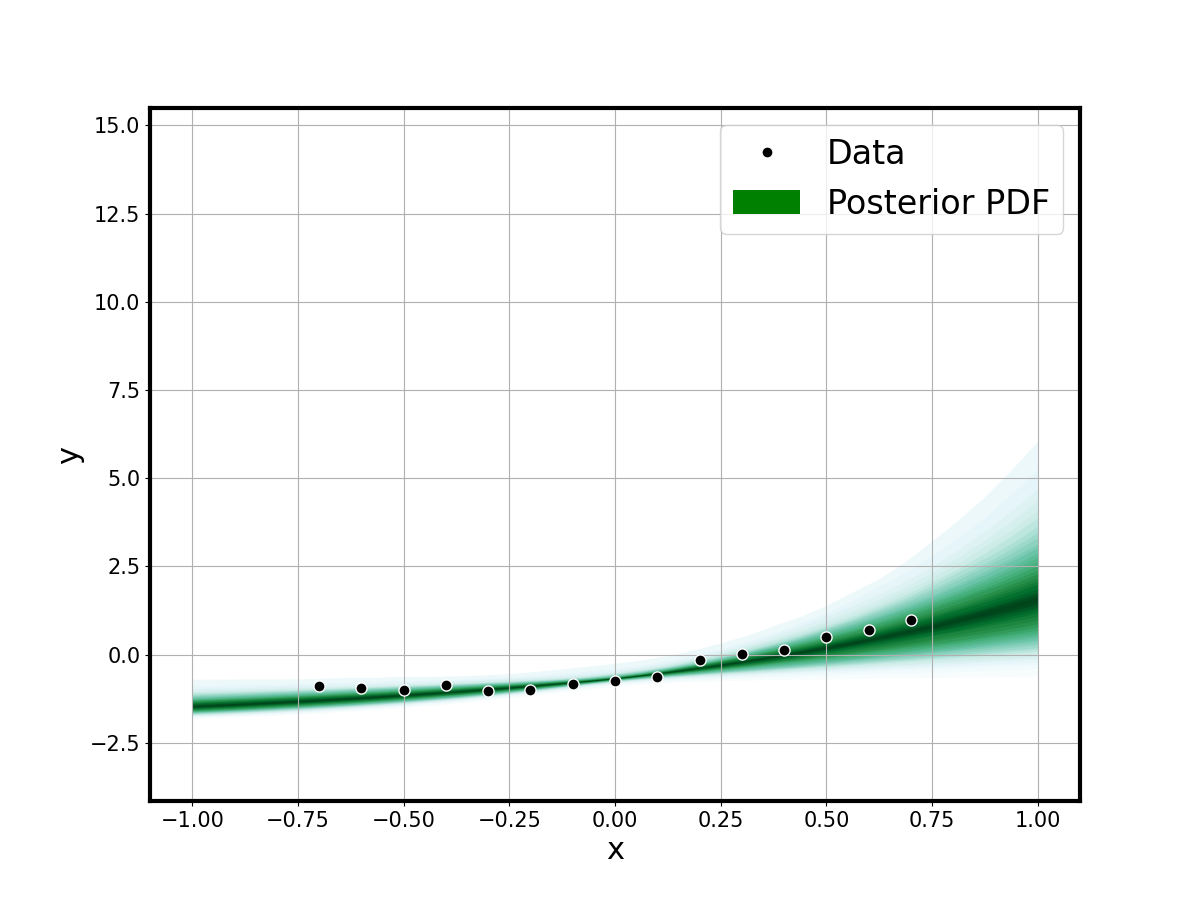
Predictive Fit
The shaded predictive envelope shows the calibrated model uncertainty against data and the true model:
fsamples_2d = np.reshape(samples['fmcmc'], (-1, ngrid), order='F')
minf.plot_1dfit_shade(xgrid, fsamples_2d.T, xydata=(xdata, ydata))
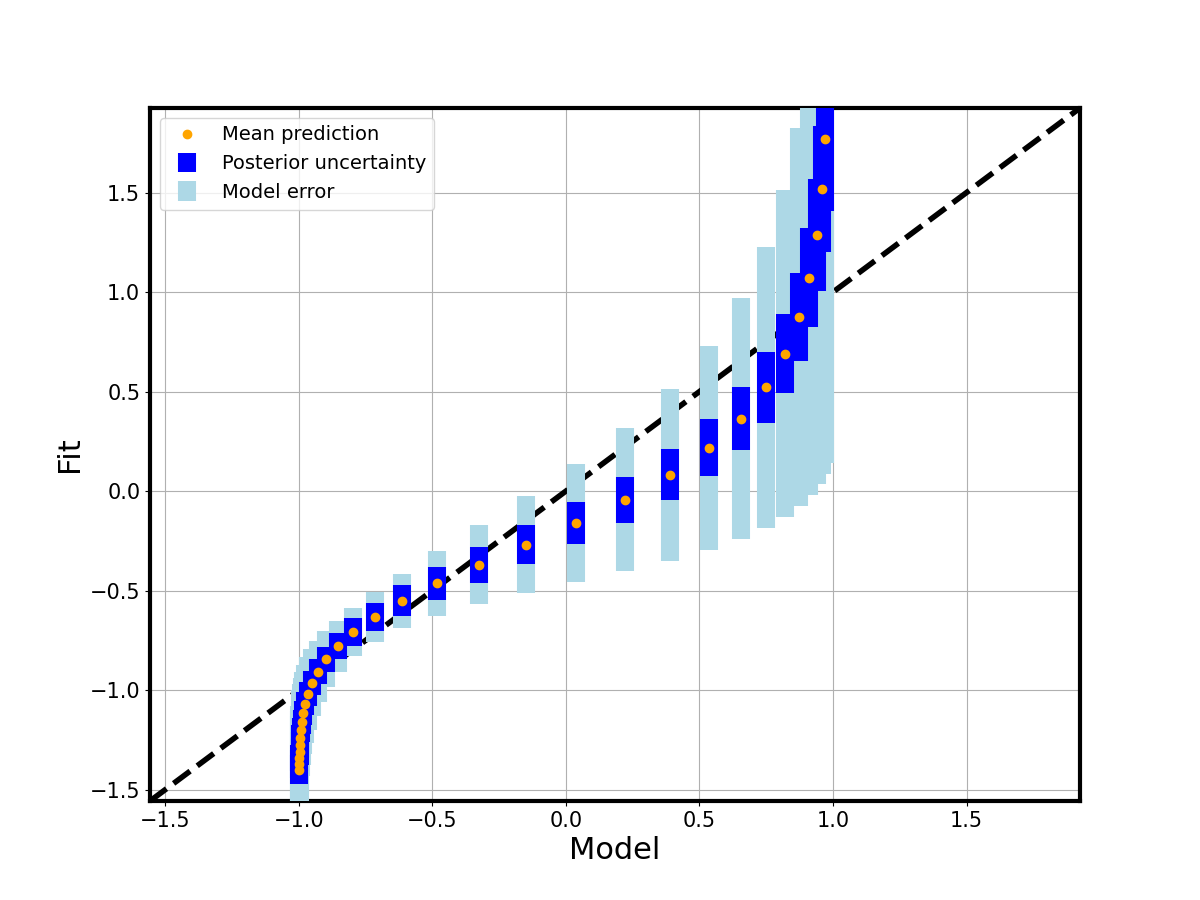
A parity plot compares the true model output per grid point with the calibrated posterior mean:
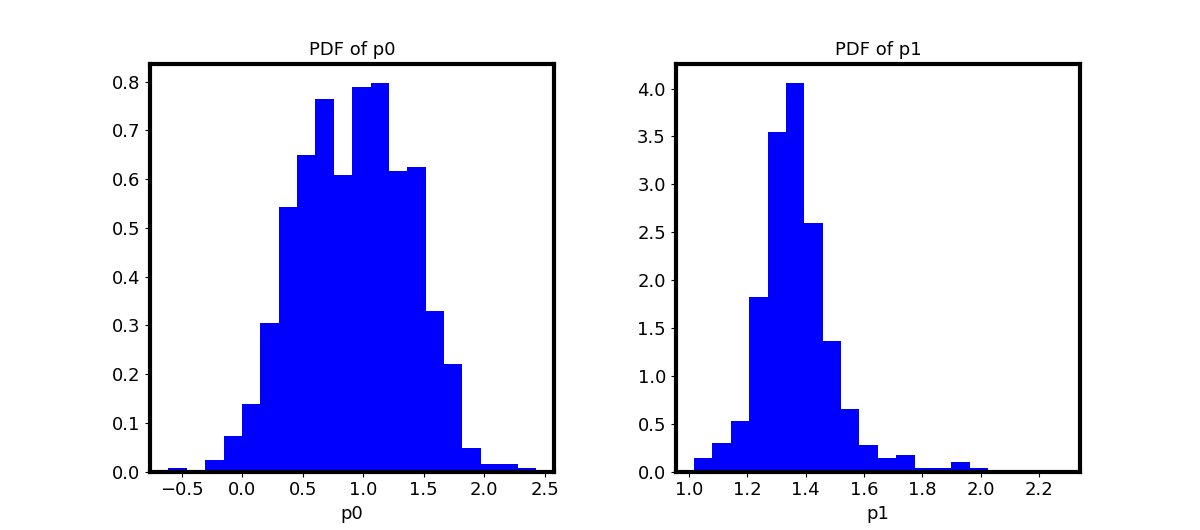
Parameter Posteriors
Marginal and joint posterior distributions of the two exponential-model parameters are visualised:
pl.plot_pdfs(samples_=psamples_2d, plot_type='inds')